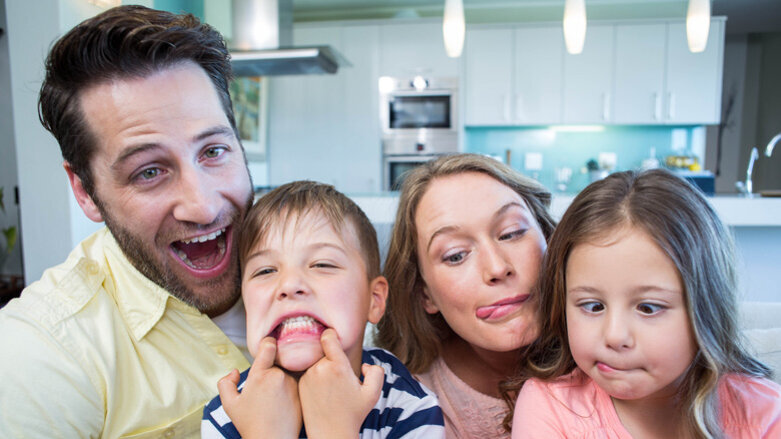
LONDON, UK: The role of the household may have an influence not only at the social level, but also at the microbe level. In a study conducted in the UK, researchers have discovered that early environmental influences are far more significant than human genetics in shaping the salivary microbiome, the group of organisms that determine oral and overall health.
Dr Adam P. Roberts, senior lecturer in antimicrobial chemotherapy and resistance at the Liverpool School of Tropical Medicine, pointed to periodontitis, which is associated with an altered microbiome, as a key example of how the study may be relevant to people’s oral and general health. “Once we understand the members of the microbiome that are responsible for health, our everyday behaviour could change to shift our microbiome favourably,” he said. Roberts co-led the study, which was conducted during his time at the UCL Eastman Dental Institute in London.
The study’s main objective was to discover how the salivary microbiome is established and what factors are most responsible for the mix of bacteria. With access to a unique sample set of DNA and saliva from an Ashkenazi Jewish family living in various households spread across four cities on three continents, the team asked how much of the variation seen in salivary microbiomes was due to host genetics and how much to the environment.
Owing to the family members adhering to ultra-Orthodox Judaism, they shared cultural diets and lifestyles that controlled for many confounding factors. Additionally, because the family members’ DNA had already been sequenced to the level of single changes in the DNA code, the research team had a unique and precise measurement of their genetic relatedness.
From this, UCL Genetics Institute graduate student Liam Shaw and the team of researchers sequenced the bacterial DNA signatures present in saliva samples from 157 family members and 27 unrelated Ashkenazi Jewish controls. Across all samples, they found that the core salivary microbiome was made up of bacteria from the Streptococcus, Rothia, Neisseria and Prevotella genera. “What that tells us is that the contact and sharing of microbes that goes on at the very local environment is what determines the differences between individuals,” said Shaw.
To understand what might be driving differences at the bacterial species level, Shaw and the team used statistical methods adopted from ecology to ascertain which factors were responsible for the most variation. When comparing factors such as shared household, city, age and genetic relatedness, the factor that determined who had the most similar saliva microbes was overwhelmingly shared household. Furthermore, spouses, parents and children younger than 10 living in a household together had the most similar salivary microbiomes.
According to Robert, the study shows that environments shared during upbringing play a major role in determining the community of bacteria that is established and knowing that the shared environment drives the microbiome may provide the ability to one day modulate it.
The study, titled “The human salivary microbiome is shaped by shared environment rather than genetics: Evidence from a large family of closely related individuals”, was published on 12 September in mBio, an open-access journal published by the American Society for Microbiology.
Tags:
BEIJING, China: Researchers from the Chinese Academy of Sciences in Beijing are reporting to have found a vaccine containing genetically modified ...
BERGEN, Norway: Rising consumption of sugary and acidic drinks has been found to be a key factor in the development and progression of dental erosion among ...
SANGLI, India: When it comes to harmful ingredients, herbal oral care products, which usually do not contain artificial substances such as sweeteners, ...
Live webinar
Wed. 8 April 2026
6:00 pm UTC (London)
Live webinar
Thu. 9 April 2026
6:00 pm UTC (London)
Live webinar
Thu. 9 April 2026
7:00 pm UTC (London)
Prof. Moritz Kebschull, Cat Edney
Live webinar
Fri. 10 April 2026
3:00 pm UTC (London)
Live webinar
Fri. 10 April 2026
4:00 pm UTC (London)
Dr. med. dent. Henrik-Christian Carl Hollay
Live webinar
Fri. 10 April 2026
5:00 pm UTC (London)
Prof. Dr. Ali Murat Kökat
Live webinar
Fri. 10 April 2026
5:00 pm UTC (London)



 Austria / Österreich
Austria / Österreich
 Bosnia and Herzegovina / Босна и Херцеговина
Bosnia and Herzegovina / Босна и Херцеговина
 Bulgaria / България
Bulgaria / България
 Croatia / Hrvatska
Croatia / Hrvatska
 Czech Republic & Slovakia / Česká republika & Slovensko
Czech Republic & Slovakia / Česká republika & Slovensko
 France / France
France / France
 Germany / Deutschland
Germany / Deutschland
 Greece / ΕΛΛΑΔΑ
Greece / ΕΛΛΑΔΑ
 Hungary / Hungary
Hungary / Hungary
 Italy / Italia
Italy / Italia
 Netherlands / Nederland
Netherlands / Nederland
 Nordic / Nordic
Nordic / Nordic
 Poland / Polska
Poland / Polska
 Portugal / Portugal
Portugal / Portugal
 Romania & Moldova / România & Moldova
Romania & Moldova / România & Moldova
 Slovenia / Slovenija
Slovenia / Slovenija
 Serbia & Montenegro / Србија и Црна Гора
Serbia & Montenegro / Србија и Црна Гора
 Spain / España
Spain / España
 Switzerland / Schweiz
Switzerland / Schweiz
 Turkey / Türkiye
Turkey / Türkiye
 UK & Ireland / UK & Ireland
UK & Ireland / UK & Ireland
 International / International
International / International
 Brazil / Brasil
Brazil / Brasil
 Canada / Canada
Canada / Canada
 Latin America / Latinoamérica
Latin America / Latinoamérica
 USA / USA
USA / USA
 China / 中国
China / 中国
 India / भारत गणराज्य
India / भारत गणराज्य
 Pakistan / Pākistān
Pakistan / Pākistān
 Vietnam / Việt Nam
Vietnam / Việt Nam
 ASEAN / ASEAN
ASEAN / ASEAN
 Israel / מְדִינַת יִשְׂרָאֵל
Israel / מְדִינַת יִשְׂרָאֵל
 Algeria, Morocco & Tunisia / الجزائر والمغرب وتونس
Algeria, Morocco & Tunisia / الجزائر والمغرب وتونس
 Middle East / Middle East
Middle East / Middle East































To post a reply please login or register